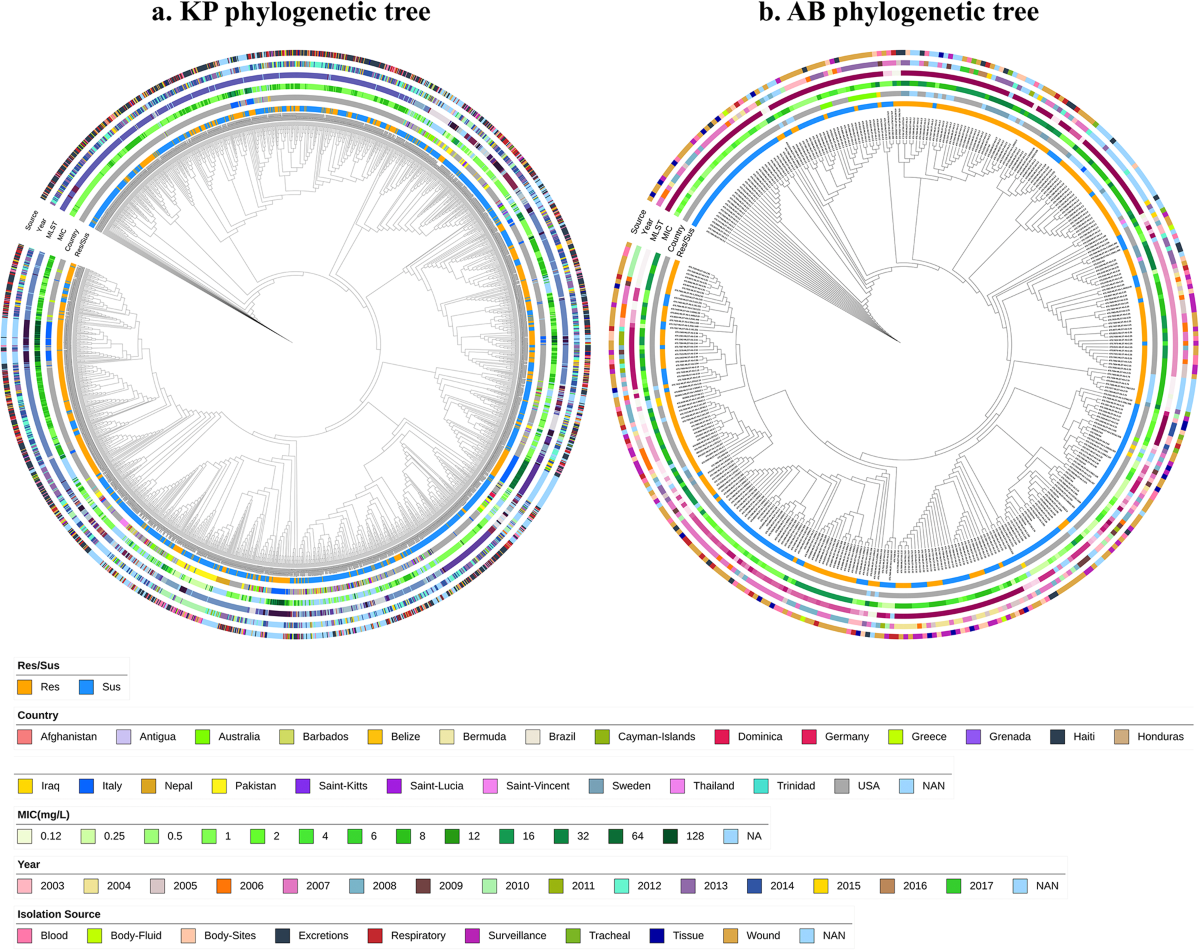
Antimicrobial resistance (AMR) poses a significant threat to global public health, with particular concern surrounding multidrug-resistant (MDR) pathogens such as Klebsiella pneumoniae and Acinetobacter baumannii. Among the antibiotics used for severe bacterial infections, meropenem, a carbapenem antibiotic, has played a vital role due to its broad-spectrum activity. However, resistance to meropenem has emerged worldwide, complicating treatment options and increasing morbidity and mortality rates. Recent advances in genome analysis, especially leveraging machine learning (ML), deep learning (DL), and association mining techniques, have enabled scientists to unravel the complex patterns and mechanisms underpinning this resistance on a global scale.
The Importance of Genome Analysis in Understanding Resistance Patterns
Traditional microbiological methods have laid the foundation for understanding AMR mechanisms, but they often fall short in deciphering the full complexity of resistance gene dissemination and interactions. Genome analysis, on the other hand, allows for an intricate view of the genetic composition of resistant strains, revealing insights into resistance determinants, mobile genetic elements, and the evolutionary trajectories that facilitate resistance spread.
By applying advanced computational tools such as ML and DL, researchers are now able to identify hidden patterns within massive genomic datasets. These techniques improve predictive accuracy regarding resistance phenotypes and help in detecting previously unnoticed associations between genes, resistance traits, and clinical factors. Association mining further complements this approach by uncovering statistically significant correlations, aiding in the identification of co-resistance patterns and potential resistance plasmids or integrons involved in the dissemination of carbapenem resistance.
Global Patterns of Meropenem Resistance in Klebsiella pneumoniae
Geographical Distribution and Resistance Trends
The genome-based studies reveal that meropenem resistance in Klebsiella pneumoniae exhibits distinct geographical patterns influenced by regional antimicrobial usage, healthcare practices, and environmental factors. Notably, resistance rates are highest in regions with widespread carbapenem use, such as parts of Asia, the Middle East, and certain areas of Europe. In contrast, countries with stringent antimicrobial stewardship programs tend to show lower resistance levels.
Genomic analyses demonstrate that resistance in Klebsiella pneumoniae primarily arises due to the acquisition of carbapenemase genes, such as blaKPC, blaNDM, blaOXA-48, and blaVIM. These genes are frequently located on mobile genetic elements like plasmids, enabling horizontal gene transfer between strains and species, thus accelerating resistance spread.
Genomic Features Driving Resistance
- Carbapenemase Production: The primary mechanism, rendered effective by the presence of genes such as blaKPC, blaNDM, and others.
- Efflux Pumps: Overexpression of efflux systems like AcrAB-TolC enhances drug expulsion mechanisms.
- Porin Loss or Modification: Mutations or deletions in porin proteins diminish antibiotic entry into bacterial cells.
ML and DL models trained on genomic features successfully predict resistance phenotypes, highlighting the most influential genetic determinants and providing a framework for rapid diagnostics.
Resistance Patterns in Acinetobacter baumannii
Emergence and Amplification of Resistance
Acinetobacter baumannii has become notorious for its ability to survive in hospital environments and its capacity to acquire multiple resistance determinants. Genome analysis shows that meropenem resistance here is often multifactorial, involving a combination of gene acquisition and intrinsic resistance mechanisms.
Common resistance determinants include carbapenemase genes such as blaOXA-23, blaOXA-24, and blaOXA-58. These are frequently embedded within large multidrug resistance (MDR) islands, facilitating co-resistance to other antibiotic classes, including aminoglycosides and fluoroquinolones.
Genetic Factors and Resistance Co-occurrence
- Class D OXA-type Carbapenemases: The dominant resistance mechanism, often associated with high-level meropenem resistance.
- Integrons and Transposons: Genetic elements that mobilize multiple resistance genes simultaneously.
- Efflux Pump Overexpression: Genes such as adeABC, contribute to resistance by actively exporting drugs out of bacterial cells.
Association mining techniques reveal that the presence of specific carbapenemase genes correlates strongly with resistance to other antibiotic classes, leading to extensively drug-resistant (XDR) and pandrug-resistant (PDR) phenotypes. This co-resistance complicates treatment regimens and demands more intensive infection control strategies.
Implications for Global Public Health and Future Directions
The integration of genome analysis with ML/DL and association mining provides a powerful platform to monitor and predict resistance trends dynamically. Such insights are crucial for designing targeted antimicrobial stewardship programs, developing molecular diagnostics for rapid detection, and informing policy decisions.
Key takeaways include:
- High prevalence of carbapenemase genes like blaKPC, blaNDM, and blaOXA across various regions, with notable regional differences.
- The significant role of mobile genetic elements in disseminating resistance traits between strains and species.
- The importance of understanding co-resistance patterns to address treatment challenges effectively.
- The need for global surveillance systems integrating genomic and epidemiological data for early detection and containment of resistant strains.
Conclusion
The application of advanced genome analysis techniques illuminates the complex landscape of meropenem resistance in critical bacterial pathogens like Klebsiella pneumoniae and Acinetobacter baumannii. Understanding global resistance patterns at the genetic level enables clinicians, microbiologists, and policymakers to craft more effective interventions, optimize treatment protocols, and ultimately curb the spread of multidrug-resistant bacteria.
As resistance continues to evolve, ongoing research utilizing machine learning, deep learning, and association mining will remain essential in combating this global health threat. Future efforts should focus on expanding genomic surveillance, fostering international collaboration, and developing new antimicrobials and diagnostics to stay ahead of resistant pathogens.
For more updated news please keep visiting Hourly Prime News.