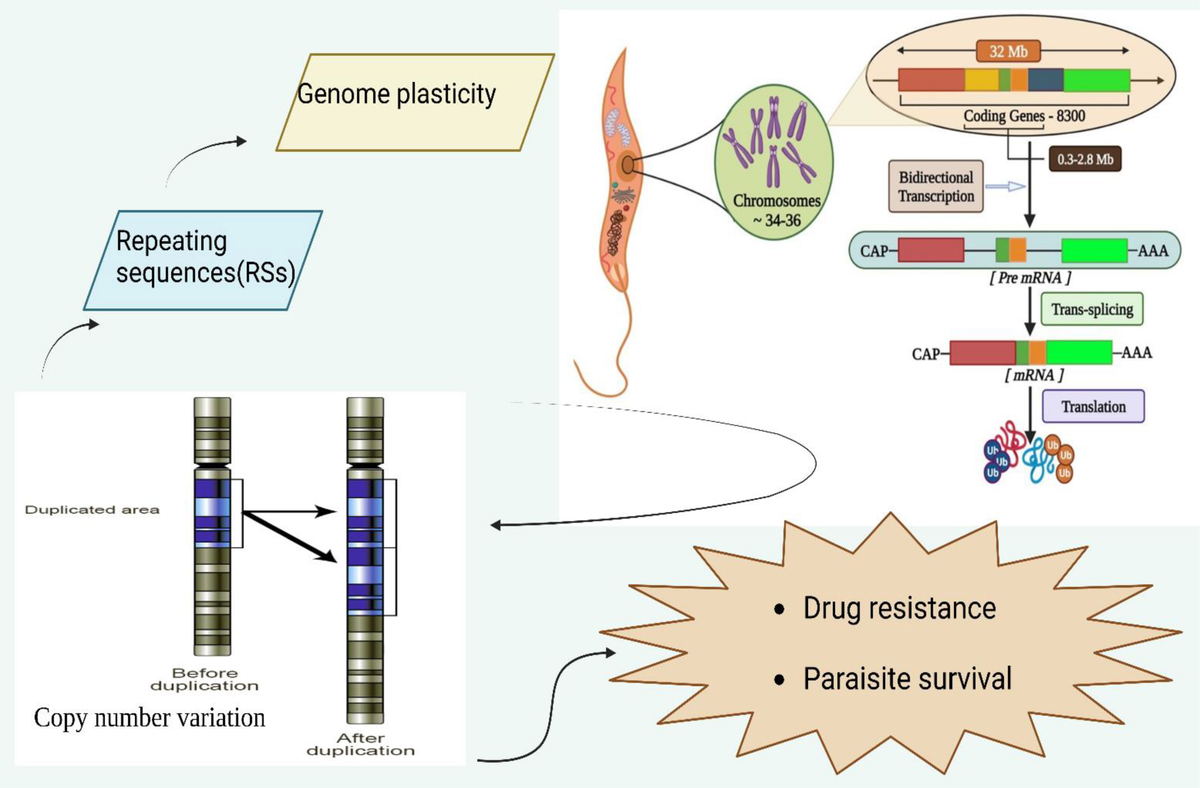
Leishmania donovani is a parasitic protozoan responsible for visceral leishmaniasis, a severe disease affecting millions worldwide. Understanding the molecular mechanisms underpinning its survival and proliferation is crucial for developing targeted treatments. Among these mechanisms, DNA replication—an essential process for cellular division—is of particular interest, especially the proteins involved and their interactions. Recent investigations have shed light on the critical role of Cdc45, a key component of the eukaryotic DNA replication machinery, and its interacting partners within Leishmania donovani. These discoveries not only enhance our knowledge of parasite biology but also open avenues for novel therapeutic interventions.
Role of Cdc45 in DNA Replication: An Overview
Cdc45 (Cell division cycle 45) is a conserved protein that functions as a pivotal factor in the initiation and progression of DNA replication. Within the context of eukaryotic cells, Cdc45 forms part of the replicative helicase complex, primarily activating the MCM (minichromosome maintenance) complex and facilitating the unwinding of DNA strands. This unwinding is essential for allowing DNA polymerases to replicate the genetic material accurately.
In *Leishmania donovani*, like in many other organisms, Cdc45 is a part of the replication fork machinery, coordinating with other proteins such as MCM helicase complexes and the GINS complex. Given the parasitic nature and unique biology of Leishmania, studying how Cdc45 interacts with specific proteins in this organism provides insights into its replication process and potential vulnerabilities.
Identification of Cdc45-Interacting Partners in Leishmania donovani
Recent groundbreaking research has employed advanced proteomic techniques to identify the interaction partners of Cdc45 in Leishmania donovani. A notable study published in BMC Microbiology used affinity purification coupled with mass spectrometry to map out the protein interaction network centered around Cdc45.
**Key Findings:**
- Two novel proteins involved in DNA replication were identified as Cdc45 interactors:
- These proteins appear to be exclusive to *Leishmania donovani* and are absent in humans, making them attractive targets for drug development.
This discovery underscores the complexity of the parasite’s replication machinery and hints at specialized adaptions that could be exploited therapeutically.
Significance of These Proteins
The identified proteins are believed to participate directly in DNA unwinding and replication fork progression. Their interaction with Cdc45 suggests a cooperative role in forming a replication complex tailored to Leishmania’s unique cellular processes.
By characterizing these proteins’ structures and functions, scientists can better understand how version-specific variations in the replication machinery contribute to the parasite’s survival and pathogenicity. These insights are instrumental, especially considering the challenge of drug resistance in current leishmaniasis treatments.
Emergence of New Druggable Targets in Leishmania
Complementing the findings about Cdc45 interaction partners, recent research from the Max-Planck-Gesellschaft has reported the discovery of a novel protein in Leishmania parasites that is considered *druggable*. This protein appears to be pivotal for parasite viability and represents a promising candidate for targeted therapy.
**Main Highlights:**
- The recently identified protein plays a critical role in DNA replication and cell cycle regulation.
- Its unique features and essential functions make it an excellent candidate for therapeutic intervention, especially since it has no human homologs.
This discovery is particularly significant because it offers a new pathway for designing drugs that can selectively target the parasite without affecting human cells.
Implications for Therapeutic Strategies
The combined insights from Cdc45 interaction studies and the newly discovered druggable proteins present an exciting frontier in anti-leishmanial drug development. Targeting vital proteins involved in DNA replication can potentially halt parasite proliferation, contributing to more effective treatments with fewer side effects.
Furthermore, the specificity of these proteins to Leishmania species minimizes the risk of off-target effects, promising more precise therapeutic interventions. Future research aimed at high-throughput screening and structural analysis of these proteins will accelerate the development of potent inhibitors.
Concluding Remarks: Paving the Way Forward
The recent revelations of key proteins associated with Cdc45 and the identification of novel druggable proteins underscore a pivotal shift in our understanding of Leishmania donovani’s molecular biology. By dissecting the intricacies of its DNA replication machinery, researchers have unveiled vulnerabilities that could revolutionize treatment strategies for leishmaniasis.
As science advances, integrative approaches combining proteomics, molecular biology, and drug design will be essential. Targeting parasite-specific replication proteins offers a promising route toward effective, selective therapeutics, ultimately aiding millions affected by this neglected tropical disease.
**In summary:**
– The identification of Cdc45-interacting partners has expanded our understanding of Leishmania donovani’s DNA replication machinery.
– Two novel proteins involved in replication processes have been uncovered, offering potential therapeutic targets.
– The discovery of a new druggable protein specific to Leishmania underscores the potential for highly selective anti-parasitic drugs.
– Continued research into these proteins and their interactions is critical for developing next-generation treatments.
By unraveling the complex interplay of proteins in Leishmania’s replication machinery, scientists move closer to designing targeted drugs that could significantly improve disease management and patient outcomes.
For more updated news please keep visiting Hourly Prime News.